Label-efficient deep learning for kidney pathology
This work package pursues a number of questions relating to the implementation of deep-learning models towards achieving computer-assisted diagnostics in nephropathology.
Background:
Using traditional approaches, examining a kidney biopsy in order to establish a disease diagnosis can be a cumbersome task involving numerous repetitive steps, such as the detection and classification of structural lesions.
Aims and objectives:
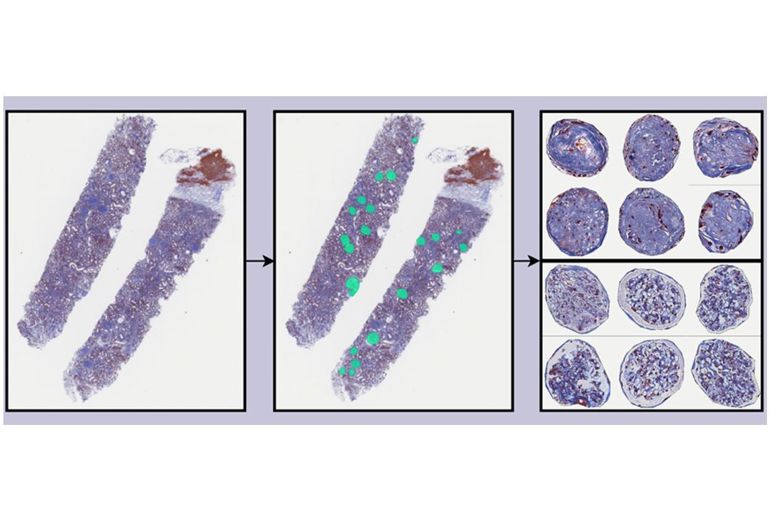
In order to assist pathologists in their routine diagnostic duties, the project seeks to explore how deep-learning might be used to automate key steps in the diagnostic process. Specifically, a major objective pertains to the automatic characterization of glomeruli, which are the basic filtering units of the kidney and are frequently affected by disease-related morphological changes. For this purpose, several deep-learning strategies will be evaluated for e.g. the detection, segmentation, clustering, and classification of such structures from biopsy images.
Additional objectives include:
- the general prediction of disease classes, diagnosis, clinical endpoints/variables, and possible therapy options from biopsy images,
- approaching explainable AI-solutions for better understanding/utilizing the morphological features employed by deep learning models, and
- and addressing the problem of obtaining sufficient data for training.
Specifically, establishing a deep-learning model typically requires large amounts of labeled data, but annotated kidney biopsy images are hard to come by and are only scarcely available in the literature. Thus, the project also seeks to establish and publish a related database of images to aid the community, and to explore more label-efficient (unsupervised, semi-supervised, weakly supervised, active learning) approaches to training deep neural networks.
Sist oppdatert 11.01.2024